- Research Briefing
- Published:
Subjects
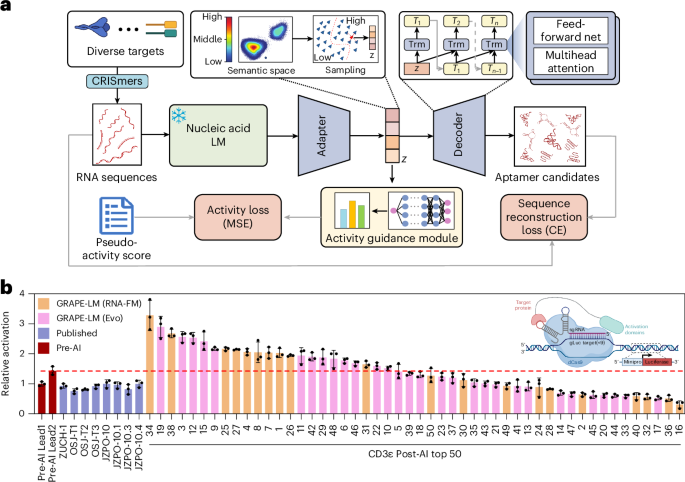
GRAPE-LM (generator of RNA aptamers powered by activity-guided evolution and language model) is a generative AI framework that enables the one-round generation of short RNA binders. When guided by CRISPR–Cas-based intracellular screening, GRAPE-LM outperforms traditional multi-round methods for aptamer evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout

References
-
Du, Z. et al. Molecular bioengineering of functional nucleic acids. Nat. Rev. Bioeng. https://doi.org/10.1038/s44222-025-00361-y (2025). A review article that presents the functional properties of nucleic acids, with a focus on engineering, optimization and biomedical prospects.
-
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990). This paper reports the technology of systematic evolution of ligands by exponential enrichment (SELEX) for RNA aptamer selection.
-
Zhang, J., Lang, M., Zhou, Y. & Zhang, Y. Predicting RNA structures and functions by artificial intelligence. Trends Genet. 40, 94–107 (2024). A review article that presents recent advances in AI-driven techniques for predicting RNA structure and function and for RNA design, offering directions and insights for future studies on the sequence–structure–function relationships of RNAs.
-
Iwano, N., Adachi, T., Aoki, K., Nakamura, Y. & Hamada, M. Generative aptamer discovery using RaptGen. Nat. Comput. Sci. 2, 378–386 (2022). This paper reports a method that uses a variational autoencoder (VAE) framework with a profile hidden Markov model and SELEX data to design nucleic acid aptamers.
-
Zhang, J. et al. Repurposing CRISPR/Cas to discover SARS-CoV-2 detecting and neutralizing aptamers. Adv. Sci. 10, 2300656 (2023). This paper introduces CRISmers, a CRISPR–Cas-based screening system for RNA aptamers that targets a chosen protein of interest in an intracellular context.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This is a summary of: Zhang, J. et al. Single-round evolution of RNA aptamers with GRAPE-LM. Nat. Biotechnol. https://doi.org/10.1038/s41587-026-03007-5 (2026).
Rights and permissions
About this article
Cite this article
Generative AI powered by nucleic acid language model enables one-round evolution of RNA aptamers. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-026-03008-4
-
Published:
-
Version of record:
-
DOI: https://doi.org/10.1038/s41587-026-03008-4