- Research Briefing
- Published:
Subjects
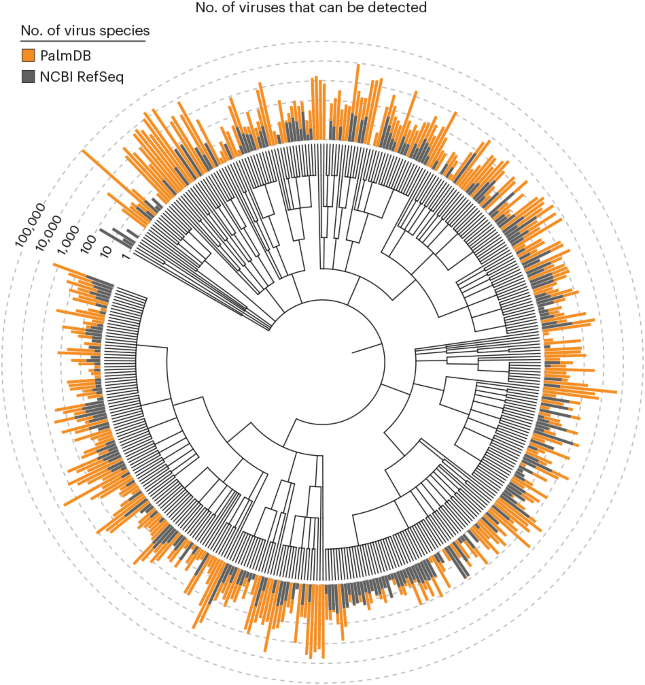
We introduce a method that detects viral sequences in RNA sequencing data on the basis of highly conserved proteins, enabling the detection of more than 100,000 RNA virus species. We analyzed the presence of novel viruses and host gene expression in parallel to characterize viral tropism and host immune responses in individual cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

References
-
Edgar, R. C. et al. Petabase-scale sequence alignment catalyses viral discovery. Nature 602, 142–147 (2022). This paper describes the creation of PalmDB.
-
Sullivan, D. K. et al. kallisto, bustools and kb-python for quantifying bulk, single-cell and single-nucleus RNA-seq. Nat. Protoc. 20, 587–607 (2024). This paper describes the RNA sequencing data alignment tools kallisto, bustools and kb-python.
-
Melsted, P. et al. Modular, efficient and constant-memory single-cell RNA-seq preprocessing. Nat. Biotechnol. 39, 813–818 (2021). This paper describes the processing of single-cell RNA sequencing data with kallisto.
-
Kotliar, D. et al. Single-cell profiling of Ebola virus disease in vivo reveals viral and host dynamics. Cell 183, 1383–1401.e19 (2020). This paper describes the creation and original analysis of the single-cell RNA sequencing data from rhesus macaques infected with Ebola virus.
-
Crick, F. H., Griffith, J. S. & Orgel, L. E. Codes Without Commas. Proc. Natl Acad. Sci. USA. 43, 416–421 (1957). In this paper, Crick et al. hypothesize that our genetic code is a comma-free code because it “gives the ‘magic number’ 20 [amino acids]”.
-
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024). This paper introduces version 6 of iTOL.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This is a summary of: Luebbert, L. et al. Detection of viral sequences at single-cell resolution identifies novel viruses associated with host gene expression changes. Nat. Biotechnol. https://doi.org/10.1038/s41587-025-02614-y (2025).
Rights and permissions
About this article
Cite this article
Agnostic viral detection at single-cell resolution reveals novel viruses. Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02638-4
-
Published:
-
DOI: https://doi.org/10.1038/s41587-025-02638-4