Data availability
The primary data supporting the findings of this study are accessible within the paper and Supplementary Information. Additional data can be obtained from the corresponding author upon reasonable request. Source data are provided with this paper.
References
-
Chater, K. F. Streptomyces inside-out: a new perspective on the bacteria that provide us with antibiotics. Philos. Trans. R. Soc. Lond. B Biol. Sci. 361, 761–768 (2006).
-
Hutchings, M. I., Truman, A. W. & Wilkinson, B. Antibiotics: past, present and future. Curr. Opin. Microbiol. 51, 72–80 (2019).
-
Zhang, L. & Demain, A. L. (eds) Natural Products: Drug Discovery and Therapeutic Medicine. (Humana Press, 2005).
-
Li, S. et al. Polyketide pesticides from actinomycetes. Curr. Opin. Biotechnol. 69, 299–307 (2021).
-
Wang, W. et al. Harnessing the intracellular triacylglycerols for titer improvement of polyketides in Streptomyces. Nat. Biotechnol. 38, 76–83 (2020).
-
Yan, H., Li, S. & Wang, W. Reprogramming naturally evolved switches for Streptomyces chassis development. Trends Biotechnol. 43, 12–15 (2025).
-
Zhang, Y. X. et al. Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415, 644–646 (2002).
-
Li, J. et al. A non-carboxylative route for the efficient synthesis of central metabolite malonyl-CoA and its derived products. Nat. Catal. 7, 361–374 (2024).
-
Thaker, M. N. et al. Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat. Biotechnol. 31, 922–927 (2013).
-
Barka, E. A. et al. Taxonomy, physiology, and natural products of actinobacteria. Microbiol. Mol. Biol. Rev. 80, 1–43 (2016).
-
Nett, M., Ikeda, H. & Moore, B. S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 26, 1362–1384 (2009).
-
Miethke, M. et al. Towards the sustainable discovery and development of new antibiotics. Nat. Rev. Chem. 5, 726–749 (2021).
-
Montano Lopez, J., Duran, L. & Avalos, J. L. Physiological limitations and opportunities in microbial metabolic engineering. Nat. Rev. Microbiol. 20, 35–48 (2022).
-
Wang, W. et al. Angucyclines as signals modulate the behaviors of Streptomyces coelicolor. Proc. Natl Acad. Sci. USA 111, 5688–5693 (2014).
-
van Wezel, G. P. & McDowall, K. J. The regulation of the secondary metabolism of Streptomyces: new links and experimental advances. Nat. Prod. Rep. 28, 1311–1333 (2011).
-
Li, S., Li, Z., Pang, S., Xiang, W. & Wang, W. Coordinating precursor supply for pharmaceutical polyketide production in Streptomyces. Curr. Opin. Biotechnol. 69, 26–34 (2021).
-
Qiu, S. et al. Building a highly efficient Streptomyces super-chassis for secondary metabolite production by reprogramming naturally-evolved multifaceted shifts. Metab. Eng. 81, 210–226 (2023).
-
Wu, J. et al. Developing a pathway-independent and full-autonomous global resource allocation strategy to dynamically switching phenotypic states. Nat. Commun. 11, 5521 (2020).
-
Gupta, A., Reizman, I. M., Reisch, C. R. & Prather, K. L. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat. Biotechnol. 35, 273–279 (2017).
-
Din, M. O. et al. Synchronized cycles of bacterial lysis for in vivo delivery. Nature 536, 81–85 (2016).
-
Alnahhas, R. N. et al. Majority sensing in synthetic microbial consortia. Nat. Commun. 11, 3659 (2020).
-
Polkade, A. V., Mantri, S. S., Patwekar, U. J. & Jangid, K. Quorum sensing: an under-explored phenomenon in the phylum actinobacteria. Front. Microbiol. 7, 131 (2016).
-
Biarnes-Carrera, M., Breitling, R. & Takano, E. Butyrolactone signalling circuits for synthetic biology. Curr. Opin. Chem. Biol. 28, 91–98 (2015).
-
Zhou, S. et al. Molecular basis for control of antibiotic production by a bacterial hormone. Nature 590, 463–467 (2021).
-
Takano, E., Chakraburtty, R., Nihira, T., Yamada, Y. & Bibb, M. J. A complex role for the gamma-butyrolactone SCB1 in regulating antibiotic production in Streptomyces coelicolor A3(2). Mol. Microbiol. 41, 1015–1028 (2001).
-
Corre, C., Song, L., O’Rourke, S., Chater, K. F. & Challis, G. L. 2-Alkyl-4-hydroxymethylfuran-3-carboxylic acids, antibiotic production inducers discovered by Streptomyces coelicolor genome mining. Proc. Natl Acad. Sci. USA 105, 17510–17515 (2008).
-
Kitani, S. et al. Avenolide, a Streptomyces hormone controlling antibiotic production in Streptomyces avermitilis. Proc. Natl Acad. Sci. USA 108, 16410–16415 (2011).
-
Wang, W. et al. Identification of a butenolide signaling system that regulates nikkomycin biosynthesis in Streptomyces. J. Biol. Chem. 293, 20029–20040 (2018).
-
Cuthbertson, L. & Nodwell, J. R. The TetR family of regulators. Microbiol. Mol. Biol. Rev. 77, 440–475 (2013).
-
Wang, W. et al. An engineered strong promoter for Streptomycetes. Appl. Environ. Microbiol. 79, 4484–4492 (2013).
-
Bhukya, H., Bhujbalrao, R., Bitra, A. & Anand, R. Structural and functional basis of transcriptional regulation by TetR family protein CprB from S. coelicolor A3(2). Nucleic Acids Res. 42, 10122–10133 (2014).
-
Wang, J. et al. A novel role of ‘pseudo’γ-butyrolactone receptors in controlling γ-butyrolactone biosynthesis in Streptomyces. Mol. Microbiol. 82, 236–250 (2011).
-
Horbal, L., Fedorenko, V. & Luzhetskyy, A. Novel and tightly regulated resorcinol and cumate-inducible expression systems for Streptomyces and other actinobacteria. Appl. Microbiol. Biotechnol. 98, 8641–8655 (2014).
-
Wang, X., Fu, Y., Wang, M. & Niu, G. Synthetic cellobiose-inducible regulatory systems allow tight and dynamic controls of gene expression in Streptomyces. ACS Synth. Biol. 10, 1956–1965 (2021).
-
Hou, J. et al. Engineering the ultrasensitive transcription factors by fusing a modular oligomerization domain. ACS Synth. Biol. 7, 1188–1194 (2018).
-
Lou, C. et al. Synthesizing a novel genetic sequential logic circuit: a push‐on push‐off switch. Mol. Syst. Biol. 6, 350 (2010).
-
Mascher, T. Past, present, and future of extracytoplasmic function σ factors: distribution and regulatory diversity of the third pillar of bacterial signal transduction. Annu. Rev. Microbiol. 77, 625–644 (2023).
-
Seipke, R. F., Patrick, E. & Hutchings, M. I. Regulation of antimycin biosynthesis by the orphan ECF RNA polymerase sigma factor σAntA. PeerJ 2, e253 (2014).
-
Bai, C. et al. Exploiting a precise design of universal synthetic modular regulatory elements to unlock the microbial natural products in Streptomyces. Proc. Natl Acad. Sci. USA 112, 12181–12186 (2015).
-
Sun, P. et al. Spiroketal formation and modification in avermectin biosynthesis involves a dual activity of AveC. J. Am. Chem. Soc. 135, 1540–1548 (2013).
-
Kitani, S., Ikeda, H., Sakamoto, T., Noguchi, S. & Nihira, T. Characterization of a regulatory gene, aveR, for the biosynthesis of avermectin in Streptomyces avermitilis. Appl. Microbiol. Biotechnol. 82, 1089–1096 (2009).
-
Qiu, J. et al. Overexpression of the ABC transporter AvtAB increases avermectin production in Streptomyces avermitilis. Appl. Microbiol. Biotechnol. 92, 337–345 (2011).
-
Hao, Y. et al. Avermectin B1a production in Streptomyces avermitilis is enhanced by engineering aveC and precursor supply genes. Appl. Microbiol. Biotechnol. 106, 2191–2205 (2022).
-
Yang, M., Hao, Y., Liu, G. & Wen, Y. Enhancement of acyl-CoA precursor supply for increased avermectin B1a production by engineering meilingmycin polyketide synthase and key primary metabolic pathway genes. Microb. Biotechnol. 17, e14470 (2024).
-
Madduri, K. et al. Production of the antitumor drug epirubicin (4′-epidoxorubicin) and its precursor by a genetically engineered strain of Streptomyces peucetius. Nat. Biotechnol. 16, 69–74 (1998).
-
Malla, S., Niraula, N. P., Liou, K. & Sohng, J. K. Improvement in doxorubicin productivity by overexpression of regulatory genes in Streptomyces peucetius. Res. Microbiol. 161, 109–117 (2010).
-
Malla, S., Niraula, N. P., Liou, K. & Sohng, J. K. Enhancement of doxorubicin production by expression of structural sugar biosynthesis and glycosyltransferase genes in Streptomyces peucetius. J. Biosci. Bioeng. 108, 92–98 (2009).
-
Scotti, C. & Hutchinson, C. R. Enhanced antibiotic production by manipulation of the Streptomyces peucetius dnrH and dnmT genes involved in doxorubicin (adriamycin) biosynthesis. J. Bacteriol. 178, 7316–7321 (1996).
-
Song, E. et al. Proteomic approach to enhance doxorubicin production in panK-integrated Streptomyces peucetius ATCC 27952. J. Ind. Microbiol. Biotechnol. 38, 1245–1253 (2011).
-
Ryu, Y. G., Butler, M. J., Chater, K. F. & Lee, K. J. Engineering of primary carbohydrate metabolism for increased production of actinorhodin in Streptomyces coelicolor. Appl. Environ. Microbiol. 72, 7132–7139 (2006).
-
Alam, K. et al. Streptomyces: the biofactory of secondary metabolites. Front. Microbiol. 13, 968053 (2022).
-
Breitling, R. et al. Synthetic biology approaches to actinomycete strain improvement. FEMS Microbiol. Lett. 368, fnab060 (2021).
-
Breitling, R. & Takano, E. Synthetic biology of natural products. Cold Spring Harb. Perspect. Biol. 8, a023994 (2016).
-
Moser, F. et al. Genetic circuit performance under conditions relevant for industrial bioreactors. ACS Synth. Biol. 1, 555–564 (2012).
-
Trosset, J. Y. & Carbonell, P. Synergistic synthetic biology: units in concert. Front. Bioeng. Biotechnol. 1, 11 (2013).
-
Xia, P. F., Ling, H., Foo, J. L. & Chang, M. W. Synthetic genetic circuits for programmable biological functionalities. Biotechnol. Adv. 37, 107393 (2019).
-
Kieser, T., Bibb, M. J., Buttner, M. J., Chater, K. F. & Hopwood, D. A. Practical Streptomyces Genetics (The John Innes Foundation, 2000).
-
Wang, W. X. et al. Identification of a butenolide signaling system that regulates nikkomycin biosynthesis in Streptomyces. J. Biol. Chem. 293, 20029–20040 (2018).
-
Rodriguez-Garcia, A., Combes, P., Perez-Redondo, R., Smith, M. C. & Smith, M. C.Natural and synthetic tetracycline-inducible promoters for use in the antibiotic-producing bacteria Streptomyces. Nucleic. Acids Res. 33, e87 (2005).
-
Wang, W. et al. Development of a synthetic oxytetracycline-inducible expression system for Streptomycetes using de novo characterized genetic parts. ACS Synth. Biol. 5, 765–773 (2016).
-
Blin, K., Pedersen, L. E., Weber, T. & Lee, S. Y. CRISPy-web: an online resource to design sgRNAs for CRISPR applications. Synth. Syst. Biotechnol. 1, 118–121 (2016).
-
Yan, H. et al. A rational multi-target combination strategy for synergistic improvement of non-ribosomal peptide production. Nat. Commun. 16, 1883 (2025).
-
Chen, H. H. et al. High-yield porphyrin production through metabolic engineering and biocatalysis. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02267-3 (2024).
-
Cao, J. et al. Harnessing a previously unidentified capability of bacterial allosteric transcription factors for sensing diverse small molecules in vitro. Sci. Adv. 4, eaau4602 (2018).
-
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25, 402–408 (2001).
-
Zhou, H. et al. Systematic development of a highly efficient cell factory for 5-aminolevulinic acid production. Trends Biotechnol. 42, 1479–1502 (2024).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (W2411016 and 32170095 to W.W. and 32121005 and 32327801 to L.Z.), the National Key Research and Development Program of China (2020YFA0907800 to L.Z. and 2022YFC2105400 to C.Z.), the Agricultural Science and Technology Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-CSCB-202401 to S.L.), the Youth Innovation Promotion Association CAS (Y202027 to W.W.) and the 111 Project (B18022 to L.Z). We would like to thank Z. Fan, G. Ai, E. Li and T. Zhao from IMCAS for the BLI assay, LC–MS/MS analysis, UPLC–MS analysis and flow cytometry assay, respectively.
Ethics declarations
Competing interests
W.W., B.Y. and Z.L. have filed provisional patents for this work to the China National Intellectual Property Administration (CN 202510569133.9 and CN 202510569134.3). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Shuguang Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DNA alignment of identified binding sites of the promoters regulated by quorum sensing receptors.
The sequences were aligned using MUSCLE, the conserved palindromic sequences (identity > 70%) were highlighted in yellow, the completely conserved DNA bases were marked with asterisks.
Extended Data Fig. 2 Determination of interaction between Ptrig and receptors in vitro.
a, Purification of known quorum sensing signalling molecule receptors. The purity was confirmed by SDS-PAGE analysis. Lane 1: His6-ScbR (24.9 kDa), lane 2: His6-AvaR1 (26.9 kDa), lane 3: His6-SabR1 (25.5 kDa), lane 4: His6-MmfR (25.1 kDa). b, Purification of putative quorum sensing signaling molecule receptors. The purity of the putative receptors was also confirmed by SDS-PAGE analysis. Lane 1: His6-WP_067441803.1 (24.4 kDa), Lane 2: His6-WP_205368416.1 (26.7 kDa), lane 3: His6-WP_185944307.1 (26.2 kDa), lane 4: His6-WP_012999107.1 (24.3 kDa), lane 5: His6-WP_242709678.1 (26.9 kDa), lane 6: His6-WP_226048631.1 (23.5 kDa). c, d, e and f, Interaction between Ptrig and representative receptors determined by EMSA. KasO* intergenic region (97 bp) was used as negative control (lane 1). The concentration of both kasO* and Ptrig DNA fragments used was 1 pmol. Lanes 2 to 6 show the addition of increasing amounts of receptors (0, 0.8, 1.6, 2.4, 3.2 pmol respectively) to the protein–DNA complexes. The SDS-PAGE and the EMSA assay were performed three times (n = 3; replicates are shown in Source Data files).
Extended Data Fig. 3 Determination of Ptrig response to native quorum sensing systems.
a, Temporal profiles of fluorescence intensity and intracellular MMF1 quorum sensing signaling molecule concentration in S. coelicolor A3(2) Δ scbA. b, Temporal profiles of fluorescence in quorum sensing mutant strains Δ scbR and Δ scbA, respectively. Values are shown as mean ± s.d. from three (n = 3) independent biological replicates.
Extended Data Fig. 4 Interaction between Ptrig and representative receptors from cluster V to X.
The curves were fitted by the data determined by BLI. a, WP_067441803.1 in cluster V. b, WP_205368416.1 in cluster VI. c, WP_185944307.1 in cluster VII. d, WP_012999107.1 in cluster VIII. e, WP_242709678.1 in cluster IX. f, WP_226048631.1 in cluster X.
Extended Data Fig. 5 Characterization and optimization of repression systems.
a, Determination of leakage in four inducible systems by evaluating the fluorescence without inducers. b, Evaluation of the toggle effect of bistable genetic circuit. Medium with cumate or cellulose was refreshed every 18 h. c, Temporal variation of fluorescence without inducers in the strain harboring unoptimized bistable circuit. d, Schematic illustration of the tetramer design of the repressors. e, Schematic of genetic circuits used for evaluating the regulatory behavior of the optimized repressors. The output promoter activity was indicated by fluorescence of sfgfp reporter, while the expression of the optimized repressor was driven by the PTAC promoter which is induced by IPTG. The input promoter activity was measured by the same sfgfp gene, and the IPTG inducer was employed to induce the input promoter via deactivating to the LacI repressor. f, and g, Dose−response curves of the native dimer and artificial tetramer CymR*/CebR*. h, Evaluation of four variants of PcebR. i, Evaluation of the toggle effect of redesigned bistable gene circuit. Medium with cumate or cellulose was changed every 18 h. For a, b, c, f, g, h and i, values are shown as mean ± s.d. from three (n = 3) independent biological replicates.
Extended Data Fig. 6 Performance of stabilizer and amplifier module on transcriptional level.
a, Quantification of gene expression outputs from strains with or without packaged stabilizer module. The expression levels were measured by RT-qPCR. Relative expression values at 12 hours were normalized to a value of one. b, Characterization the amplification effect by comparing the relative transcriptional levels in the strains with and without amplifier module. c, Comparison of the GFP fluorescence intensity between S. venezuelae wild type (left) and engineering strain harboring sfgfp driven by PON mutants (right) using flow cytometry. d, Fluorescence activated cell sorting to obtain S. venezuelae with varying strength of PON mutants. Gating strategy was based on GFP fluorescence intensity. e, Correlation between transcriptional and translational profiles of 10 selected mutant promoters derived from PON. The transcriptional profile was evaluated by quantitative analysis of sfgfp driven by these promoters, while translational efficiency was evaluated based on the fluorescence intensity of sfGFP. For a and b, values are shown as mean ± s.d. from three (n = 3) independent biological replicates.
Extended Data Fig. 7 Comparison of native quorum sensing system and artificial control system SMARTS.
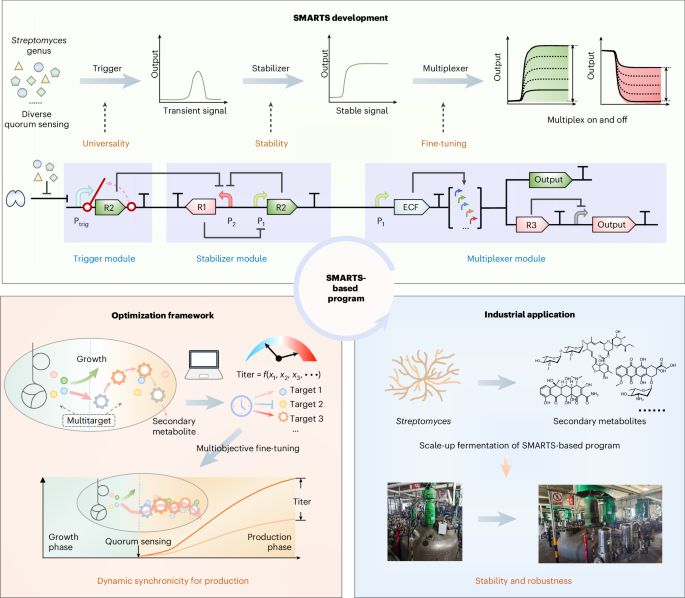
A quorum sensing signal transduction pathway in Streptomyces griseus was selected as a representative to elucidate the native regulatory mechanism. This signal transduction pathway can be divided into three stages: recognition, transduction and output. The native quorum sensing system was adapted for dynamic physiological regulation in Streptomyces species. In contrast, by integrating trigger, stabilizer and multiplexer module, the artificial control system SMARTS can response to diverse quorum sensing, and consequently convert the transient quorum sensing signals into stable, multiplexed outputs with adjustable strength in different Streptomyces strains. Furthermore, after responding to the quorum sensing signal, SMARTS is independently regulated and demonstrates complete orthogonality with the endogenous regulatory system.
Extended Data Fig. 8 Baiweimectin activity against soil parasitic nematodes.
a, Survival curve of nematodes treated with baiweimectin compared to commercially available fosthiazate. b, LC50 and LC90 values derived from the survival curve. For a, values are shown as mean ± s.d. from four (n = 4) independent biological replicates.
Extended Data Fig. 9 Central composite design of baiweimectin optimization targets.
a, aveR. b, avtAB. c, meiC. d, fadD. e, ecaA–bicA. f, gRNA targeting sucCD. Values are shown as mean ± s.d. from four (n = 3) independent biological replicates. The statistical analysis is based on two-tailed unpaired Student’s t-test.
Extended Data Fig. 10 Optimization the production of epidoxorubicin.
a, Group I targets for enhancing epidoxorubicin production. Sources and the corresponding reactions of the targets were annotated. DOX, doxorubicin; Epi, epidoxorubicin; ε-RHO, ε-rhodomycinone; RHOD, rhodomycin D. b, Group II targets for enhancing epidoxorubicin production with annotated sources and corresponding enzymatic reactions. Ac-CoA, acetyl-CoA; M-CoA, malonyl-CoA bldD. c-i, Central composite design of epidoxorubicin optimization targets. c, dnrIN. d, avtE. e, dnrSQ. f, gRNA targeting dnrH. g, panK. h, acc. i, bldD. For c-i, values are shown as mean ± s.d. from four (n = 3) independent biological replicates. The statistical analysis is based on two-tailed unpaired Student’s t-test.
Supplementary information
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, B., Li, Z., Zhang, J. et al. Scalable secondary metabolite production in Streptomyces using a plug-and-play system. Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02762-1
-
Received:
-
Accepted:
-
Published:
-
DOI: https://doi.org/10.1038/s41587-025-02762-1